- Peptide-based antimicrobials

Associate Professor
Florentine Marx-Ladurner
phone: +43/512/9003-70207
email: florentine.marx[at]i-med.ac.at
Group members
CV
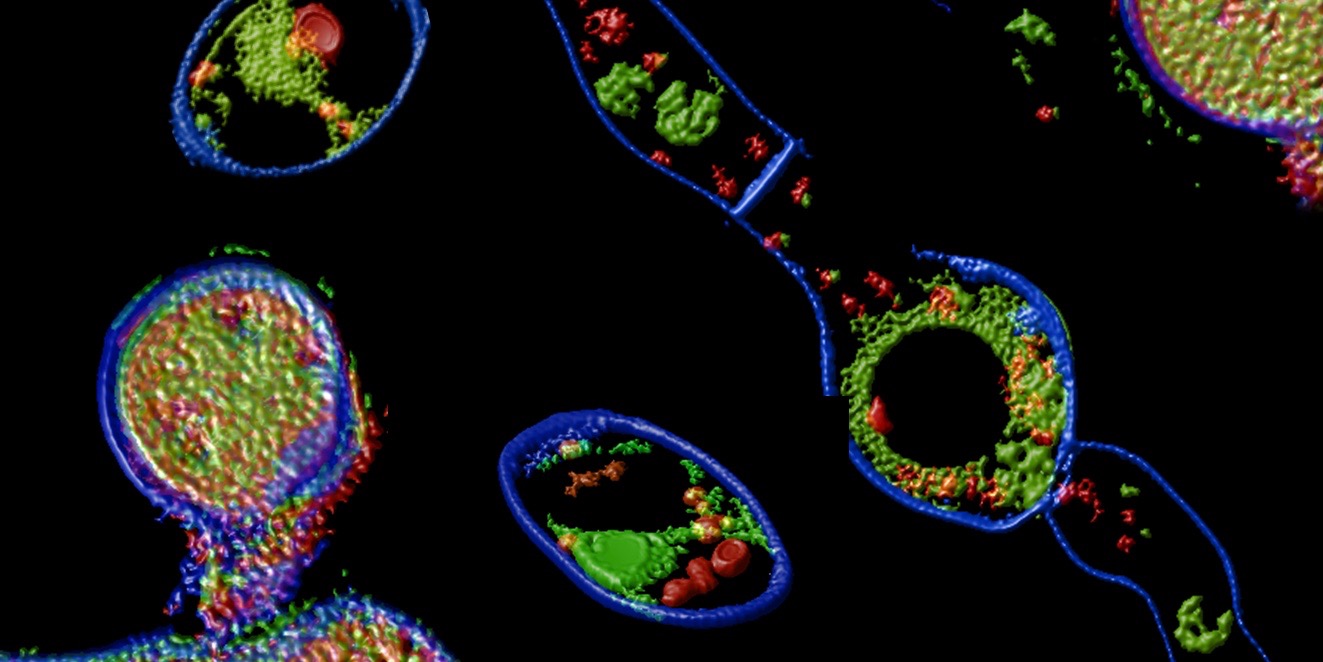
Nature-inspired antimicrobial peptides and proteins
The rise in drug resistance among human-pathogenic bacteria and fungi has created an urgent need to identify alternative antimicrobial agents with novel modes of action that differ from existing ones and impede the development of resistance. Promising candidates include small, cationic antimicrobial proteins and peptides (APs), which are natural defence molecules produced by organisms across all phyla. These APs serve as powerful model molecules for developing new and effective antimicrobial strategies for therapeutic applications.
Our main scientific interest is to characterize novel APs from diverse natural sources at the structural, functional, and molecular levels. Understanding their antimicrobial modes of action is a crucial prerequisite for the development of novel therapeutics.
Selected Publications
Marx F., Haas H., Reindl M., Stoeffler G., Lottspeich F. and Redl B. (1995) Cloning, structural organization and regulation of expression of the Penicillium chrysogenum paf gene encoding an abundantly secreted protein with antifungal activity. Gene 167,167-171.
Kaiserer L., Oberparleiter C., Weiler-Goerz R., Burgstaller W., Leiter E., Marx F. (2003) Characterization of the Penicillium chrysogenum antifungal protein PAF. Arch. Microbiol. 180: 204-210.
Oberparleiter C., Kaiserer L., Haas H., Ladurner P., Andratsch M., Marx F. (2003) Active internalization of the Penicillium chryosgenum antifungal protein PAF in sensitive aspergilli. Antimicrob. Agents Chemother. 47: 3598-3601.
Marx F. (2004) Small, basic antifungal proteins secreted from filamentous asocmycetes: a comparative study regarding expression, structure, function and potential application. Review. App. Microbiol. Biotechnol. 65: 133-142.
Leiter E., Marx F., Pusztahelyi T., Haas H., Pocsi I. (2004) Penicillium chrysogenum glucose oxidase – a study on its antifungal effects, J. Appl. Microbiol. 97: 1201-1209.
Marx F., Salvenmoser W., Kaiserer L., Grässle S., Weiler-Görz R., Zadra I., Oberparleiter C. (2005) Proper folding of the antifungal protein PAF is required for optimal protein activity. Res. Microbiol. 156: 35-46.
Theis T., Marx F., Salvenmoser W., Stahl U., Meyer W. (2005) New insights into the target site and mode of action of the antifungal protein (AFP) of Aspergillus giganteus. Res. Microbiol. 156: 47-56.
Galgoczy, L., Papp, T., Leiter, E., Marx, F., Pocsi, I., Vagvolgyi, C. (2005) Sensitivity of different zygomycetes to the Penicillium chrysogenum antifungal protein (PAF). J. Basic Microbiol. 45: 136-141.
Sappanos, H., Szigeti, G.P., Pal, B., Rusznak, Z., Szucs, G., Rajnavolgyi, E., Balla, J., Balla, G., Nagy, E., Leiter, E., Pocsi, I., Marx, F., Csernoch, L. (2005) The Penicillium chrysogenum-derived antifungal peptide shows no toxic effects on mammalian cells in the intended therapeutic concentration. Naunyn Schmiedeberg’s Arch. Pharmacol. 371: 122-132.
Leiter, E., Szappanos, H., Oberparleiter, C., Kaiserer, L., Csernoch, L., Pusztahelyi, T., Emri, T., Pocsi, I., Salvenmoser, W., Marx, F. (2005) Antifungal protein PAF severely affects the integrity of the plasma membrane of Aspergillus nidulans and induces an apoptosis-like phenotype. Antimicrob. Agents Chemother. 49: 2445-2453.
Hagen, S., Marx, F., Ram, A.R., and Meyer, V. (2007). The antifungal protein AFP from Aspergillus giganteus inhibits chitin synthesis in sensitive fungi. Appl. Environm. Microbiol. 73, 2128-2134.
Marx, F., Binder, U., Leiter, E., and Pocsi, I. (2008) The Penicillium chrysogenum antifungal protein PAF, a promising tool for fungal cell biology studies and the development of new antifungal therapies. Cell. Mol. Life Sci. 65, 445-454.
Batta, G., Barna, T., Gaspari, Z., Sandor, S., Köver, K., Binder, U., Sarg, B., Kaiserer, L., Chhillar, A., Eigentler, A., Leiter, E., Hegedüs, N., Pocsi, I., Lindner, H., and Marx, F. (2009) Functional aspects of the solution structure and dynamics of PAF – a highly stable antifungal protein from Penicillium chrysogenum. FEBS J. 276(10):2875-2890
Binder, U., Oberparleiter, C., Meyer V., and Marx F. (2010) The antifungal protein PAF interferes with PKC/MPK and cAMP/PKA signalling of Aspergillus nidulans Mol Microbiol. 75: 294-307.
Binder, U., Chu, M., Read, N.D., and Marx, F. The antifungal activity of the Penicillium chrysogenum protein PAF disrupts calcium homeostasis in Neurospora crassa. (2010). Eukaryot. Cell. 9: 1374-1382.
Hegedüs, N., Sigl, C., Zadra, I., Pócsi, I., and Marx, F. The paf gene product modulates asexual development in Penicillium chrysogenum. (2011). J. Basic Microbiol. 51: 1-10.
Hegedüs, N., Leiter, E., Kovacs, B., Tomori, V., Kwon, NJ., Emri, T., Marx, F., Batta, G., Csernoch, L., Haas, H., Yu, JH.; Pocsi, I. (2011) The small molecular mass antifungal protein of Penicillium chrysogenum – a mechanism of action oriented review. J. Basic Microbiol. 51(6); 561-571.
Hegedüs, N., and Marx, F. (2013). Antifungal proteins: More than antimicrobials? Fungal Biol. Rev. 26: 132-145.
Fizil, Á., Gáspári, Z., Barna, T., Marx, F., and Batta, G. (2015). „Invisible“ conformers of an antifungal disulfide protein revealed by constrained cold an heat unfolding, CEST-NMR experiments, and molecular dynamics calculations. Chemistry 21: 5136-5144.
Binder, U., Bencina, M., Fizil, Á., Batta, G., Chhillar, A.K., and Marx, F. (2015). Protein kinase A signaling and calcium ions are major players in PAF mediated toxicity against Aspergillus niger. FEBS Lett. 589: 1266-1271.
Virágh, M., Marton, A., Vizler, C., Tóth, L., Vágvölgyi, C., Marx, F., and Galgóczy L. (2015). Insight into the antifungal mechanism of Neosartorya fischeri antifungal protein. Protein Cell. 6: 518-528.
Sonderegger, C., Galgóczy, L., Garrigues, S., Fizil, Á., Borics, A., Manzanares, P., Hegedüs, N., Huber, A., Marcos, JF. Batta, G., and Marx, F. (2016) A Penicillium chrysogenum-based expression system for the prodcution of small, cysteine-rich antifungal proteins for structural and functional analyses. Microb Cell Fact. 15:192.
Sonderegger C., Fizil Á., Burtscher L., Hajdu D., Muñoz A., Gáspári Z., Read ND., Batta G., Marx F. (2017). D19S Mutation of the Cationic, Cysteine-Rich Protein PAF: Novel Insights into Its Structural Dynamics, Thermal Unfolding and Antifungal Function. PLoS ONE 12(1):e0169920.
Garrigues S., Gandia M., Borics A., Marx F., Manzanares P., Marcos J.F. (2017). Mapping and identification of antifungal peptides in the putative antifungal protein AfpB from the filamentous fungus Penicillium digitatum. Frontiers Microbiol. 8: 592.
Gágóczy L., Borics A., Virágh M., Ficze H., Váradi G., Kele Z., Marx F. (2017). Structural determinants of Neosartorya fischeri antifungal protein (NFAP) for folding, stability and antifungal activity. Sci. Reports 7:1963.
Garrigues S., Gandía M., Popa C., Borics A., Marx F., Coca M., Marcos J., Manzanares P. (2017). Efficient production and characterization of the novel and highly active antifungal protein AfpB from Penicillium digitatum. Sci. Reports 7: 14663.
Huber A, Hajdu D., Bratschun-Khan D., Gáspári Z., Varbanov M., Philippot S., Fizil Á., Czajlik A., Kele Z., Sonderegger C., Galgóczy L., Bodor A., Marx F., Batta G. (2018). New antimicrobial potential and structural properties of PAFB: a cationic, cysteine-rich protein from Penicillium chrysogenum Q176. Sci. Reports 8: 1751.
Tóth L., Váradi G., , Borics A., Batta G., Kele Z., Vendrinszky Á., Tóth R., Ficze H., Tóth G.K., Vágvölgyi C., Marx F., Galgóczy, L. (2018). Anti-candidal activity and functional mapping of recombinant and synthetic Neosartorya fischeri antifungal protein 2 (NFAP2). Front. Microbiol. 9: 393.
Sonderegger C., Váradi G., Galgóczy L., Kocsubé S., Posch W., Borics A., Dubrac S., Tóth G.K., Wilflingseder D., Marx F. (2018). The evolutionary conserved g-core motif influences the anti-Candida activity of the Penicillium chrysogenum antifungal protein PAF. Front. Microbiol. 9: 1655.
Garrigues S., Gandía M., Castillo L., Coca M., Marx F., Marcos J.F., Manzanares P. (2018). Three antifungal proteins from Penicillium expansum: Different patterns of production and antifungal activity. Front. Microbiol 9: 2370.
Fizil Á., Sonderegger C., Czajlik A., Fekete A., Komáromi I., Hajdu D., Marx F., Batta G. (2018). Calcium binding of the antifungal protein PAF: Structure, dynamics and function aspects by NMR and MD simulations. PLoS One 13: e0204825.
Kovács R, Holzknecht J, Hargitai Z, Papp C, Farkas A, Borics A, Tóth L, Váradi G, Tóth GK, Kovács I, Dubrac S, Majoros L, Marx F, Galgóczy L. (2019). In vivo applicability of Neosartorya fischeri antifungal protein 2 (nfap2) in treatment of vulvovaginal candidiasis. Antimicrob. Agents Chemother.63(2): e01777-18.
Hajdu D., Huber A., Czajlik A., Tóth L., Kele Z., Kocsubé S., Fizil Á., Marx F., Galgóczy L., Batta G. (2019). Solution structure and novel insights into phylogeny and mode of action of the Neosartorya (Aspergillus) fischeri antifungal protein (NFAP). Int. J. Biol. Macromol. 129:511-522
Galgóczy L, Marx F. (2019). Do antimicrobial proteins contribute to overcoming the hidden antifungal crisis at the dawn of a post-antibiotic era? Microorganisms 7(1).
Huber A, Oemer G, Malanovic N, Lohner K, Kovács L, Salvenmoser W, Zschocke J, Keller MA, Marx F. (2019) Membrane Sphingolipids Regulate the Fitness and Antifungal Protein Susceptibility of Neurospora crassa. Front. Microbiol. 10: 605
Galgóczy L, Yap A, Marx F. (2019) Cysteine-Rich Antifungal Proteins from Filamentous Fungi are Promising Bioactive Natural Compounds in Anti-Candida Therapy. Isr. J. Chem. 59(5):360-370. Review
Huber A, Lerchster H, Marx F. (2019) Nutrient Excess Triggers the Expression of the penicillium chrysogenum Antifungal Protein PAFB. Microorganisms 7 (12): 654
Tóth L, Boros É, Poór P, Ördög A, Kele Z, Váradi G, Holzknecht J, Bratschun-Khan D, Nagy I, Tóth GK, Rákhely G, Marx F, Galgóczy L. (2020). The potential use of the Penicillium chrysogenum antifungal protein PAF, the designed variant PAFopt and its γ-core peptide Pγopt in plant protection. Microb. Biotechnol. 13(5):1403-1414 Epub ahead of pring.
Tóth L, Váradi G, Boros É, Borics A, Ficze H, Nagy I, Tóth GK, Rákhely G, Marx F, Galgóczy L. (2020) Biofungicidal Potential of Neosartorya (Aspergillus) Fischeri Antifungal Protein NFAP and Novel Synthetic gamma-Core Peptides. Front Microbiol. 11:820
Huber A, Galgóczy L, Váradi G, Holzknecht J, Kakar A, Malanovic N, Leber R, Koch J, Keller MA, Batta G, Tóth GK, Marx F. (2020) Two small, cysteine-rich and cationic antifungal proteins from Penicillium chrysogenum: A comparative study of PAF and PAFB. Biochim Biophys Acta Biomembr. 1862(8):183246.
Holzknecht J, Kühbacher A, Papp C, Farkas A, Váradi G, Marcos JF, Manzanares P, Tóth GK, Galgóczy L, Marx F. (2020) The Penicillium chrysogenum Q176 Antimicrobial Protein PAFC Effectively Inhibits the Growth of the Opportunistic Human Pathogen Candida albicans. J Fungi (Basel). 6(3):141.
Czajlik A, Holzknecht J, Galgóczy L, Tóth L, Poór P, Ördög A, Váradi G, Kühbacher A, Borics A, Tóth GK, Marx F, Batta G. (2021) Solution Structure, Dynamics, and New Antifungal Aspects of the Cysteine-Rich Miniprotein PAFC. Int J Mol Sci. https://doi.org/10.3390/ijms22031183.
Guanini F, Huber A, Alex JM, Marx F, Crowley PB. (2021) Porous Assembly of an Antifungal Protein Mediated by Zinc and Sulfonato-calix[8]arene. J Struct Biol. https://doi.org/10.1016/j.jsb.2021.107711
Gandía, M., Kakar, A., Giner-Llorca, M., Holzknecht, J., Martínez-Culebras, P., Galgóczy, L., Marx, F., Marcos, J.F., and Manzanares, P. (2021) Potential of antifungal proteins (AFPs) to control Penicillium postharvest fruit decay. J. Fungi, 7, 449. https://doi.org/10.3390/jof7060449
Kakar, A., Holzknecht, J., Dubrac, S., Gelmi, M.L., Romanelli, A., and Marx, F. (2021) New perspectives in the antimicrobial activity of the amphibian Temporin B: Peptide analogs are effective inhibitors of Candida albicans growth. J. Fungi, 7, 457. https://doi.org/10.3390/ jof7060457
Tóth L, Poór P, Ördög A, Váradi G, Farkas A, Papp C, Bende G, Tóth GK, Rákhely G, Marx F, and Galgóczy L. (2022). The combination of Neosartorya (Aspergillus) fischeri antifungal proteins with rationally designed γ-core peptide derivatives is effective for plant and crop protection.. Biocontrol (Dordr). 67(2):249-262. doi: 10.1007/s10526-022-10132-y.
Holzknecht J., Dubrac S., Hedtrich S., Galgóczy L., and Marx F. (2022). Small, Cationic Antifungal Proteins from Filamentous Fungi Inhibit Candida albicans Growth in 3D Skin Infection Models. Microbiol Spectr 10(3). e0029922. doi: 10.1128/spectrum.00299-22.
Kakar A., Sastré-Velásquez L.E., Hess M., Galgóczy L., Papp C., Holzknecht J., Romanelli A., Váradi G., Malanovic N., Marx F. (2022). The Membrane Activity of the Amphibian Temporin B Peptide Analog TB_KKG6K Sheds Light on the Mechanism That Kills Candida albicans. mSphere. 7(5):e0029022. doi: 10.1128/msphere.00290-22.
To D., Kakar A., Kali G., Wibel R., Knoll P., Marx F., Bernkop-Schnürch A. (2023). Iminated aminoglycosides in self-emulsifying drug delivery systems: Dual approach to break down the microbial defense. J Colloid Interface Sci. 630(Pt B):164-178. doi: 10.1016/j.jcis.2022.10.077.
Váradi G., Batta G., Galgóczy L., Hajdu D., Fizil À., Czajlik A., Virágh M., Kele Z., Meyer V., Jung, S., Marx F., Tóth G. (2023). Confirmation of the disulfide connectivity and strategies of chemical synthesis of the four-disulfide-bond-stabilized Aspergillus giganteus antifungal protein, AFP. J Nat Sci. 86:782-790. doi: 10.1021/acs.jnatprod.2c00954.
Akkus-Dagdeviren Z.B., Saleh A., Schöpf C., Truszkowska M., Bratschun-Khan D., Fürst A., Seybold A., Offterdinger M., Marx F., Bernkop-Schnürch A. (2023). Phosphatase-degradable nanoparticles: A game-changing approach for the delivery of antifungal proteins. J Coll Interface Sci 646:290-300. doi: 10.1016/j.jcis.2023.05.051.
Holzknecht J, Marx F. Navigating the fungal battlefield: cysteine-rich antifungal proteins and peptides from Eurotiales. Front Fungal Biol 2024, 5:1451455. doi: 10.3389/ffunb.2024.1451455.
Truszkowska M, Stengel D, Schmidt MR, Marx F, Coraca-Huber D, Bernkop-Schürch A. Synergistic antimicrobial effect of daptomycin and ethyl lauroyl arginate containing self-emulsifying drug delivery system against bacterial infections. J Drug Deliv Sci Technol 2024, 102:A. doi:10.1016/j.jddst.2024.106324.
Bende G, Zsindely N, Laczi K, Kristóffy Z, Papp C, Farkas A, Tóth L, Sáringer S, Bodai L, Rákhely G, Marx F, Galgóczy L. The Neosartorya (Aspergillus) fischeri antifungal protein NFAP2 has low potential to trigger resistance development in Candida albicans in vitro. Microbiol Spectr 2025 13:e0127324. doi: 10.1128/spectrum.01273-24.
Gai J, File M, Erdei R, Czajlik A, Marx F, Galgóczy L, Váradi G, Batta G. Small disulfide proteins with antifungal impact: NMR experimental structures as compared to models of alphafold versions. Int J Mol Sci 2025 26:1247. doi: 10.3390/ijms26031247.
Schöpf C, Knapp M, Scheler J, Coraça-Huber DC, Romanelli A, Ladurner P, Seybold AC, Binder U, Würzner R, Marx F. The antibacterial activity and therapeutic potential of the amphibian-derived peptide TB_KKG6K. mSpere 2025 10:e01016-24. doi: 10.1128/msphere.01016-24.
Schöpf C, Geschindt A, Knapp M, Seybold AC, Coraça-Huber DC, Ausserlechner MJ, Romanelli A, Marx F. Amphibian-derived peptide analog TB_KKG6K: A powerful drug candidate against Candida albicans with anti-biofilm efficacy. bioRxiv 2025 preprint. doi: 10.1101/2025.08.22.671737
Schöpf C, Marx F. 3D human epithelial models: a promising platform for studying Candida infections and novel antifungal therapeutic options. Front Cell Infect Microbiol 2025 15:1680261. doi: 10.3389/fcimb.2025.1680261
Schöpf C, Geschindt A, Knapp M, Seybold AC, Coraça-Huber DC, Ausserlechner MJ, Romanelli A, Marx F. Amphibian-derived peptide analog TB_KKG6K: A powerful drug candidate against Candida albicans with anti-biofilm efficacy. J. Fungi, 12, 11. doi: 10.3390/jof12010011
Blanco Massani M, To D, Gintsburg D, Molnar L, Polidori I, Meile S, Seybold A, Ünalan B, Coraca-Huber DC, Marx F, Loessner MJ, Schmelcher M, Zapotoczny S, Bernkop-Schnürch A. Phage endolysin-polyphosphate/alginate nanocomplexes inhibit staphylococcal biofilms for implant protection. Carbohydr Polym Tech Appl 13, 101089. doi: 10.1016/j.carpta.2026.101089



