Bioinformatics
Research Focus
General Facts
Research
Selected Publications
Selected Funding, Collaboration
Keywords: Bioinformatics, computational biology, cancer immunology, colorectal cancer
Research (ÖSTAT Classification) : 102004, 106005
Research Focus
The research of the Institute of Bioinformatics focuses on cancer immunology. Our aim is to decipher tumour-immune cell interaction by means of a combined computational-experimental approach. We specifically look at the question of how the immune system shapes the mutation spectrum of the tumour, both during progression and after treatment, in patients with colorectal cancer. We use tumour-derived organoids, perturbation experiments and multimodal analysis, including next-generation sequencing, phosphoproteomics and imaging mass cytometry to facilitate precision immuno-oncology.
General Facts
The Institute for Bioinformatics was established in 2010 as the latest member of the Biocentre at the Medical University of Innsbruck and it is dedicated to supporting basic research and teaching in bioinformatics and computational biology. The Institute for Bioinformatics is well equipped both for data processing and management as well as for computational analysis of a broad range of data types, including next-generation sequencing data, proteomics and metabolomics data, and image analysis. It has a state-of-the-art computational infrastructure, which is dedicated to bioinformatics analysis. A number of analytical pipelines, databases and software tools developed by the laboratory and others are installed and continuously maintained (see http://icbi.at).
Research
The long-term research goal of the Institute for Bioinformatics is to establish a mechanistic rationale for immunotherapy-based combination regimens and to develop a precision immuno-oncology platform for colorectal cancer. The concept integrates a living biobank of patient-derived tumour organoids with high-throughput and high-content data for testing drug combinations as well as machine learning, in order to make therapeutic recommendations for individual patients. We are investigating three immunomodulatory processes and their respective drug classes: 1) immunostimulatory effects of selected chemotherapeutics; 2) rewiring of signalling networks induced by targeted drugs and their interference with immunity; 3) metabolic reprogramming of T-cells to enhance anti-tumour immunity and its modulation by means of drugs.
For the first time, recent advancements in molecular profiling techniques, the development of organoid technology and novel computational methods are providing the opportunity to tackle this challenging problem. Our systems-level dissection of cancer-immune cell interaction will facilitate the development of a recommendation algorithm to inform precision immuno-oncology in CRC and form the basis for an iterative platform to identify effective therapeutic regimens for individual patients.
As we have previously shown, the immune contexture – i.e. the density, composition, location and functional orientation of TILs – has a major prognostic value in CRC patients [1]. Clinical observations supported by experimental evidence with mouse models indicate that chemotherapeutics as well as targeted drugs modulate the immune contexture and thus affect disease outcome (reviewed in [2]). A provocative hypothesis was postulated, that the efficacy of most conventional and targeted cancer drugs approved for use in humans not only involves direct cytostatic/cytotoxic effects but also relies on the reactivation of tumour-targeting immune responses [2]. In accordance with this hypothesis, the goal of cancer therapy should consist in reinstating immunological control of tumour growth. It is also notable that the mechanistic rationale behind the development of cancer immunotherapy differs from the rationale underlying conventional or targeted agents, i.e. reinstating immunological control of tumour growth rather than targeting tumour cells [2]. Whilst this notion is gaining in popularity, examination of the complex interactions of tumours with the immune system has, until recently, posed considerable challenges, owing to the lack of suitable cellular and animal systems and owing to the inadequacy of the available molecular tools for comprehensive immunological characterisation.
In the past, two key technologies were developed, which allow systematic investigation of the tumour-immune interface: 1) patient-derived tumour organoids that retain cell-cell and cell-matrix interactions, which more closely resemble those of the original tumour in comparison with cells grown in two dimensions on plastic dishes; 2) high-throughput and high-content technologies, including whole exome and RNA sequencing of bulk tissue, single-cell RNA sequencing, mass-spectrometry-based proteomics [3] and multiplexed staining of tissue sections. Moreover, considerable progress has been made in the development of novel computational tools for deep mining of the data. We use these tools (Fig. 1) to investigate the impact of conventional and targeted drugs on T-cell activity, by means of archived samples, patient-derived organoids and systematic perturbation of the involved pathways, and innovative computational tools. Our aim is to classify patients by using multidimensional profiling to analyse samples, and to identify biomarkers of response to immunogenic chemotherapy and consequently patients who would benefit from combination therapy. Genomic, transcriptomic, phosphoproteomic and metabolomic characterisation of the organoids facilitates the generation of key quantitative data and will ultimately lead to identification of effective drugs and drug combinations.
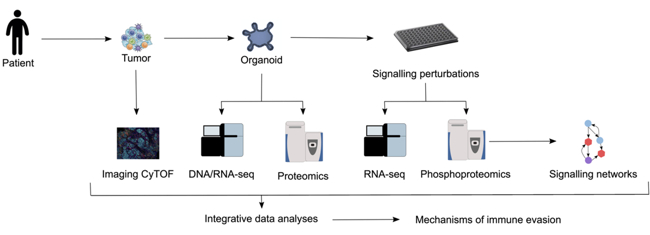
- Fridman WH, Pages F, Sautes-Fridman C, Galon J: The immune contexture in human tumours: impact on clinical outcome. Nat Rev Cancer 2012, 12:298-306.
- Galluzzi L, Buque A, Kepp O, Zitvogel L, Kroemer G: Immunological Effects of Conventional Chemotherapy and Targeted Anticancer Agents. Cancer Cell 2015, 28:690-714.
- Aebersold R, Mann M: Mass-spectrometric exploration of proteome structure and function. Nature 2016, 537:347-355.
- Dienstmann R, Vermeulen L, Guinney J, Kopetz S, Tejpar S, Tabernero J: Consensus molecular subtypes and the evolution of precision medicine in colorectal cancer. Nat Rev Cancer 2017, 17:268.
Selected Publications
- Finotello F, Rieder D, Hackl H, Trajanoski Z. Next-generation computational tools for interrogating cancer immunity. Nat Rev Genet. 20:724-746
- Finotello F, Mayer C, Plattner C, Laschober G, Rieder D, Hackl H, Krogsdam A, Loncova Z, Posch W, Wilflingseder D, Sopper S, Ijsselsteijn M, Brouwer TP, Johnson D, Xu Y, Wang Y, Sanders ME, Estrada MV, Ericsson-Gonzalez P, Charoentong P, Balko J, de Miranda N, Trajanoski Z. Molecular and pharmacological modulators of the tumor immune contexture revealed by deconvolution of RNA-seq data. Genome Med. 11:34
- Efremova M, Rieder D, Klepsch V, Charoentong P, Finotello F, Hackl H, Hermann-Kleiter N, Löwer M, Baier G, Krogsdam AM, Trajanoski Z. Targeting immune checkpoints potentiates immunoediting and changes the dynamics of tumor evolution. Nat Commun. 2018. 9:32
- Hackl H, Charoentong P, Finotello F, Trajanoski Z. Computational genomics tools for dissecting tumor-immune cell interactions. Nat Rev Genet. 2016. 17:441-458
Selection of Funding
- ERC advanced grant EPIC: Enabling immuno-oncology in colorectal cancer (PI: Z. Trajanoski)
- Horizon 2020 IMI: imSAVAR:Immune safety avatar: nonclinical mimicking of the immune system effects of immunomodulatory therapies (project partner: Z Trajanoski)
- FWF/DFG: Transregio 241 (DFG) Immune-Epithelial Communication in Inflammatory Bowel Diseases (project partner: Z Trajanoski)
- FWF doc.funds Cellular Basis of Diseases: Molecular Control of Metabolism and Inflammation (project partner: Z Trajanoski)
- OeNB Anniversary Fund: Prediction of non-canonical neoantigens for next-generation colorectal cancer immunotherapy (PI: Francesca Finotello)
- OeNB Anniversary Fund: Connection of DNA damage and repair with the immunome in high-grade serous ovarian cancer (PI: Hubert Hackl)
Collaborations
- Noel de Miranda, Leiden University, Leiden, The Netherlands
- Ruedi Aebersold, ETH Zurich, Zurich, Switzerland
- Florian Greten, Henner Farin, University of Frankfurt, Frankfurt, Germany
Devices & Services
- High-performance computing and storage (https://icbi.i-med.ac.at/infrastructure.html)




